|
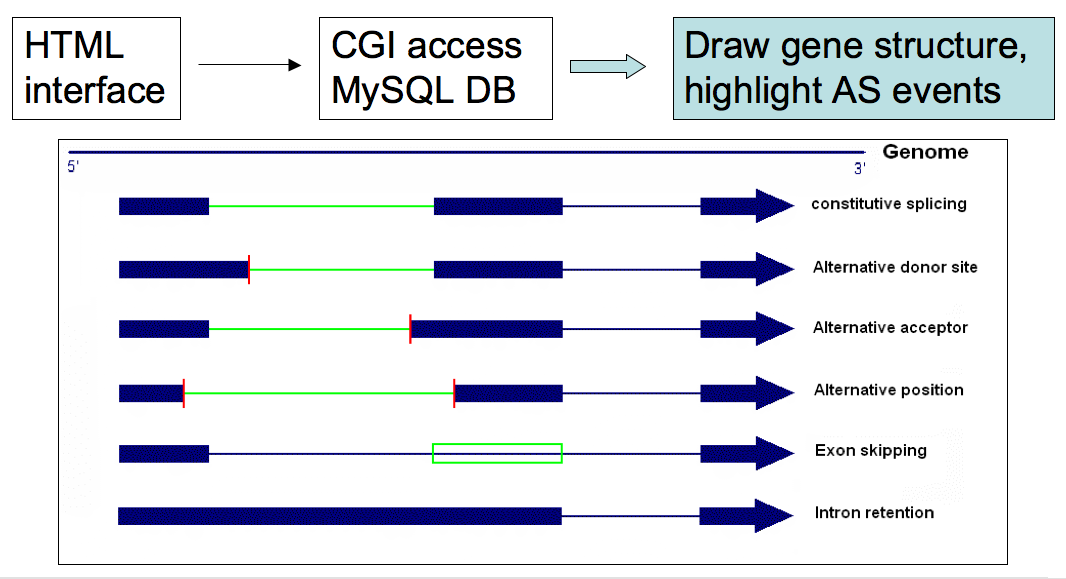
Alternative splicing visualization tool...
|
ASIP Work Flow
We identify and visualize alternative splicing events in the following three steps:
- Align all cDNA/EST sequences against their genome
using spliced-alignment program GeneSeqer. A typical
alignment was shown in the following figure:
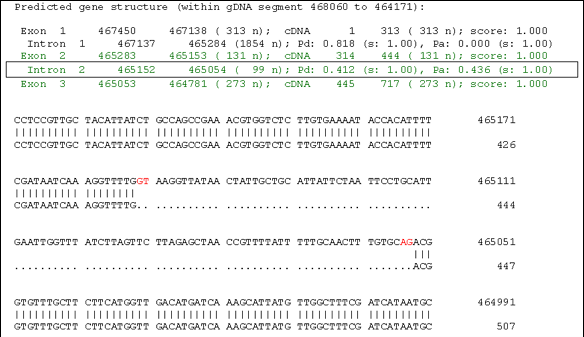
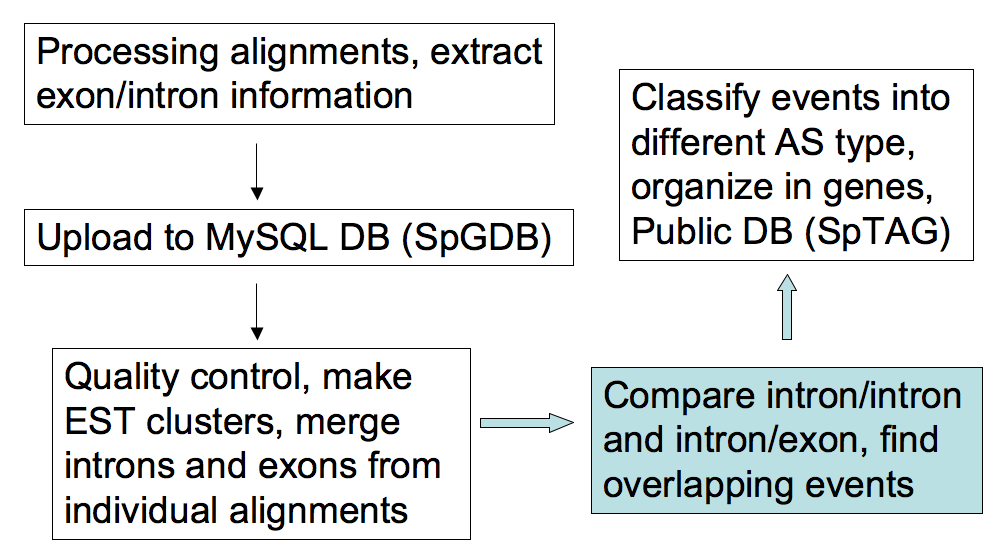
- Process the GeneSeqer alignments, extract exon/intron
information and compare their coordinates to find
overlapping introns/introns and introns/exons. Then we
classify these overlapping events into different AS type.
This process was carried out by a pipeline named ASpipe. To
ensure quality, only reliable introns and exons were
considered in this step.
- To publicize and visualize the data, we wrote an HTML
interface and used CGI script to access the database
(SpTAG). Perl module GD.pm was used to generate pictures.
AS events were specifically highlighted, as shown in the
following figure. A standalone version of these scripts
will be available soon (ASviewer)